melidosData helps you load data from the MeLiDos field study. It also contains helpers to work with REDCap dictionaries and metadata.
The project
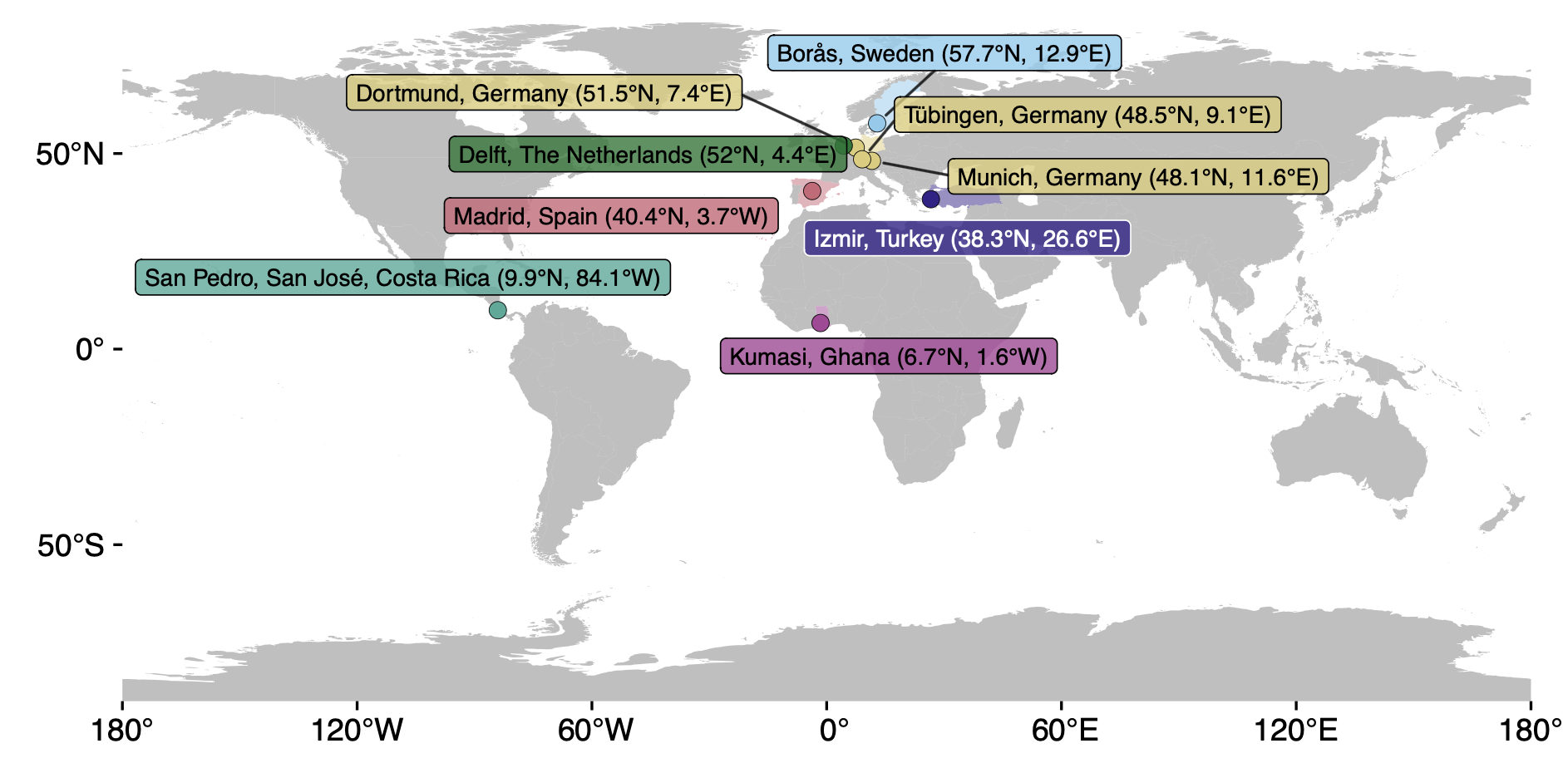
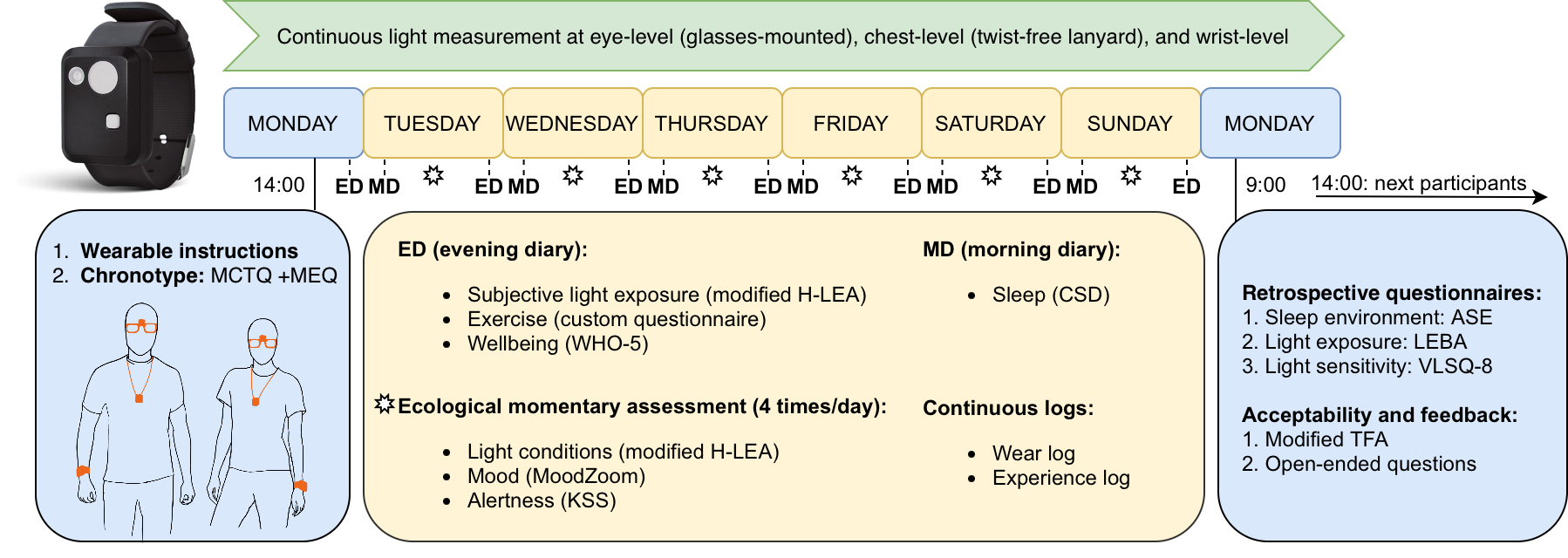
The MeLiDos field study datasets contain wearable data for personal light exposure at the eye, chest, and wrist level for 191 participants across 9 sites and 7 countries, capturing 1480 participant days of annotated data. Through a host of questionnaires at screening and discharge time, ecological momentary assessment, diaries, and logs, extensive auxiliary data are available.
- Main workflow:
-
load_data()downloads a modality for one or many sites. -
flatten_data()combines multi-site lists into one analysis-ready table, including a shared timezone.
-
| Institution (site Abbr.) | City | Country | Repository | DOI |
|---|---|---|---|---|
| RISE | Borås | Sweden | NilssonTengelinEtAl_Dataset_2026 | 10.5281/zenodo.18925834 |
| THUAS | Delft | The Netherlands | AertsEtAl_Dataset_2025 | 10.5281/zenodo.17979893 |
| BAUA | Dortmund | Germany | BroszioEtAl_Dataset_2025 | 10.5281/zenodo.18111232 |
| MPI | Tübingen | Germany | GuidolinEtAl_Dataset_2025 | 10.5281/zenodo.16895188 |
| TUM | Munich | Germany | HildenEtAl_Dataset_2025 | 10.5281/zenodo.16893901 |
| FUSPCEU | Madrid | Spain | BaezaEtAl_Dataset_2025 | 10.5281/zenodo.16834951 |
| IZTECH | Izmir | Turkey | DidikogluEtAl_Dataset_2025 | 10.5281/zenodo.16568109 |
| UCR | San José | Costa Rica | Sancho-SalasEtAl_Dataset_2025 | 10.5281/zenodo.17289456 |
| KNUST | Kumasi | Ghana | AkuffoEtAl_Dataset_2025 | 10.5281/zenodo.15576731 |
Overview of the available sites in the package
Installation
You can install the CRAN version of melidosData with:
install.packages("melidosData")You can install the development version of melidosData from GitHub with:
# install.packages("pak")
pak::pak("MeLiDosProject/melidosData")Main workflow example
library(melidosData)
library(LightLogR) #for visualization
library(dplyr) #for data manipulation
# one site -> data frame
sleep_tum <- load_data("sleepdiaries", site = "TUM")
sleep_tum |> select(Id, sleep, wake) |> head()
#> # A tibble: 6 × 3
#> Id sleep wake
#> <chr> <dttm> <dttm>
#> 1 TUM_S001 2024-05-14 00:25:00 2024-05-14 09:38:00
#> 2 TUM_S001 2024-05-15 00:17:00 2024-05-15 08:00:00
#> 3 TUM_S001 2024-05-16 00:32:00 2024-05-16 10:00:00
#> 4 TUM_S001 2024-05-17 01:25:00 2024-05-17 09:30:00
#> 5 TUM_S001 2024-05-18 00:35:00 2024-05-18 09:30:00
#> 6 TUM_S001 2024-05-19 05:32:00 2024-05-19 11:30:00
# many sites -> `melidos_data` list
sleep_all <- load_data("sleepdiaries", site = c("TUM", "UCR"))
sleep_all |> summary()
#> Length Class Mode
#> TUM 14 tbl_df list
#> UCR 14 tbl_df list
# flatten list output for analysis
sleep_all_flat <- flatten_data(sleep_all, tz = "UTC")
sleep_all_flat |>
select(site, Id, sleep, wake) |>
group_by(site) |>
slice_head(n=3)
#> # A tibble: 6 × 4
#> # Groups: site [2]
#> site Id sleep wake
#> <chr> <chr> <dttm> <dttm>
#> 1 TUM TUM_S001 2024-05-14 00:25:00 2024-05-14 09:38:00
#> 2 TUM TUM_S001 2024-05-15 00:17:00 2024-05-15 08:00:00
#> 3 TUM TUM_S001 2024-05-16 00:32:00 2024-05-16 10:00:00
#> 4 UCR UCR_S001 2025-06-16 21:35:00 2025-06-17 06:36:00
#> 5 UCR UCR_S001 2025-06-17 21:35:00 2025-06-18 06:30:00
#> 6 UCR UCR_S001 2025-06-18 22:25:00 2025-06-19 06:40:00Modalities
The following list of modalities contains the modality codes to be used in load_data().
Light exposure:
light_glasses, light_chest, light_wrist: personal light exposure datasets, captured with
ActLumusdevices, recorded in 10 second intervals for the eye-level, chest-level, and wrist-level position. Data are minimally checked after import (range, explicit gaps, device malfunctions, time shifts).light_glasses_1minute, light_chest_1minute, light_wrist_1minute: as above, but aggregated to 1-minute intervals. Furthermore, days with less than 80% of data coverage are removed. This makes the datasets both faster to download, computationally easier to work with, and also more stable to calculate light exposure metrics
Questionnaires (screening and discharge):
- health: Lifestyle and health
- demographics: Demographics
- chronotype: Chronotype (MCTQ, MEQ)
- accectability: Light glasses acceptability
- ase: Sleep environment (ASE)
- evaluation: Evaluation of study by participants
- leba: Light exposure behavior (LEBA)
- vlsq8: Light sensitivity (VLSQ-8)
Sites
The nine sites are centered on universities and research institutes. The packages contains relevant metadata for these sites:
melidos_countries
#> RISE FUSPCEU BAUA TUM
#> "Sweden" "Spain" "Germany" "Germany"
#> MPI THUAS IZTECH KNUST
#> "Germany" "The Netherlands" "Turkey" "Ghana"
#> UCR
#> "Costa Rica"
melidos_cities
#> RISE FUSPCEU BAUA
#> "Borås" "Madrid" "Dortmund"
#> TUM MPI THUAS
#> "Munich" "Tübingen" "Delft"
#> IZTECH KNUST UCR
#> "Izmir" "Kumasi" "San Pedro, San José"
melidos_colors
#> RISE FUSPCEU BAUA TUM MPI THUAS IZTECH KNUST
#> "#88CCEE" "#CC6677" "#DDCC77" "#DDCC77" "#DDCC77" "#117733" "#332288" "#AA4499"
#> UCR
#> "#44AA99"
melidos_coordinates
#> $RISE
#> [1] 57.71567 12.89087
#>
#> $FUSPCEU
#> [1] 40.41650 -3.70256
#>
#> $BAUA
#> [1] 51.498204 7.416708
#>
#> $TUM
#> [1] 48.1333 11.5667
#>
#> $MPI
#> [1] 48.5216 9.0576
#>
#> $THUAS
#> [1] 52.0116 4.3571
#>
#> $IZTECH
#> [1] 38.32 26.63
#>
#> $KNUST
#> [1] 6.675007 -1.572644
#>
#> $UCR
#> [1] 9.9372 -84.0509
melidos_tzs
#> RISE FUSPCEU BAUA
#> "Europe/Stockholm" "Europe/Madrid" "Europe/Berlin"
#> TUM MPI THUAS
#> "Europe/Berlin" "Europe/Berlin" "Europe/Amsterdam"
#> IZTECH KNUST UCR
#> "Europe/Istanbul" "Africa/Accra" "America/Costa_Rica"More information is found under the full repository of each dataset on the project page.
Mini vignette: REDCap helper workflow
# 1) load dictionary and clean labels
codebook_path <- system.file("ext", "DataDictionary_sleepdiary.csv", package = "melidosData")
codebook <- utils::read.csv(codebook_path, check.names = FALSE)
codebook_clean <- REDCap_codebook_prepare(codebook)
codebook_clean |> names()
#> [1] "Variable / Field Name"
#> [2] "Form Name"
#> [3] "Section Header"
#> [4] "Field Type"
#> [5] "Field Label"
#> [6] "Choices, Calculations, OR Slider Labels"
#> [7] "Field Note"
#> [8] "Text Validation Type OR Show Slider Number"
#> [9] "Text Validation Min"
#> [10] "Text Validation Max"
#> [11] "Identifier?"
#> [12] "Branching Logic (Show field only if...)"
#> [13] "Required Field?"
#> [14] "Custom Alignment"
#> [15] "Question Number (surveys only)"
#> [16] "Matrix Group Name"
#> [17] "Matrix Ranking?"
#> [18] "Field Annotation"
# 2) start with example REDCap export
sleep <- REDCap_example_sleep
sleep$sleepquality #original
#> [1] 3 3 2 5 2 3 3 2 4
# 3) check expected column types against dictionary metadata
check <- REDCap_coltype_check(codebook_clean, data = sleep)
check$ok
#> [1] TRUE
check$details
#> # A tibble: 16 × 6
#> col expected present actual type_ok issue
#> <chr> <chr> <lgl> <chr> <lgl> <chr>
#> 1 bedtime POSIXct TRUE POSIXct TRUE ok
#> 2 sleep POSIXct TRUE POSIXct TRUE ok
#> 3 offset POSIXct TRUE POSIXct TRUE ok
#> 4 out_ofbed POSIXct TRUE POSIXct TRUE ok
#> 5 sleepdelay numeric TRUE numeric TRUE ok
#> 6 awakenings numeric TRUE numeric TRUE ok
#> 7 awake_duration numeric TRUE numeric TRUE ok
#> 8 sleepquality numeric TRUE numeric TRUE ok
#> 9 daytype2 numeric TRUE numeric TRUE ok
#> 10 status numeric TRUE numeric TRUE ok
#> 11 scheduledate Date TRUE Date TRUE ok
#> 12 record_id character TRUE character TRUE ok
#> 13 comments character TRUE character TRUE ok
#> 14 uuid character TRUE character TRUE ok
#> 15 supplementaldata character TRUE character TRUE ok
#> 16 serializedresult character TRUE character TRUE ok
# 4) convert factor numbers into factors
sleep_factors <- REDCap_factors(sleep, codebook_clean)
sleep_factors$sleepquality #factors
#> [1] Fair Fair Poor Very good Poor Fair Fair
#> [8] Poor Good
#> Levels: Very poor Poor Fair Good Very good
# 5) attach human-readable labels from dictionary
sleep_labelled <- REDCap_col_labels(sleep_factors, codebook_clean)
sleep_labelled$sleepquality #factors with label
#> [1] Fair Fair Poor Very good Poor Fair Fair
#> [8] Poor Good
#> attr(,"label")
#> [1] How would you rate the quality of your sleep?
#> Levels: Very poor Poor Fair Good Very good
#6) add arbitrary labels, e.g. for computed variables
sleep_labelled <- sleep_labelled |> mutate(actual_sleep = sleep + sleepdelay*60)
sleep_labelled$actual_sleep |> attr("label") # incorrect label (taken from sleep)
#> [1] "What time did you try to go to sleep?"
sleep_labelled <- sleep_labelled |> add_labels(c(actual_sleep = "Time of sleep including sleepdelay (calculated)"))
sleep_labelled$actual_sleep |> attr("label") #new label
#> [1] "Time of sleep including sleepdelay (calculated)"Note: labels are volatile in R. If they are lost, e.g. after data transformation, the functions can be reexecuted.
About the MeLiDos project
MeLiDos is a joint, EURAMET-funded project involving sixteen partners across Europe, aimed at developing a metrology and a standard workflow for wearable light logger data and optical radiation dosimeters. Its primary contributions towards fostering FAIR data include the development of a common file format, robust metadata descriptors, and an accompanying open-source software ecosystem.
The project (22NRM05 MeLiDos) has received funding from the European Partnership on Metrology, co-financed from the European Union’s Horizon Europe Research and Innovation Programme and by the Participating States. Views and opinions expressed are however those of the author(s) only and do not necessarily reflect those of the European Union or EURAMET. Neither the European Union nor the granting authority can be held responsible for them.


